




















中国农业科技导报 ›› 2023, Vol. 25 ›› Issue (8): 65-75.DOI: 10.13304/j.nykjdb.2021.0876
张冬冬1( ), 韩宏伟2(
), 韩宏伟2( ), 余镇藩1, 曾斌1(
), 余镇藩1, 曾斌1( ), 杨佳惠1, 高雯雯1, 马昕彤1
), 杨佳惠1, 高雯雯1, 马昕彤1
收稿日期:2021-10-14
接受日期:2022-01-18
出版日期:2023-08-20
发布日期:2023-09-07
通讯作者:
曾斌
作者简介:张冬冬 E-mail:583787672@qq.com基金资助:
Dongdong ZHANG1( ), Hongwei HAN2(
), Hongwei HAN2( ), Zhenfan YU1, Bin ZENG1(
), Zhenfan YU1, Bin ZENG1( ), Jiahui YANG1, Wenwen GAO1, Xintong MA1
), Jiahui YANG1, Wenwen GAO1, Xintong MA1
Received:2021-10-14
Accepted:2022-01-18
Online:2023-08-20
Published:2023-09-07
Contact:
Bin ZENG
摘要:
为研究蔷薇科植物叶绿体基因组密码子偏好性,对12种蔷薇科植物进行叶绿体基因组密码子使用模式分析。结果表明,12种蔷薇科植物叶绿体基因组的GC含量为 36.54%~37.23%,密码子第3位碱基的平均GC含量为28.27%~28.61%,平均有效密码子数为47.87~49.17。筛选到32个高频密码子,主要以A/U结尾。中性绘图分析、ENC-plot分析、PR2-plot分析表明自然选择是蔷薇科植物密码子偏好性形成的主要因素。基于同义密码子相对使用度(relative synonymous codon usage,RSCU)构建的进化树与基于编码基因和叶绿体基因组构建的进化树聚类差异较大,说明基于密码子偏好特征的进化关系可能损失了一些信息。以上结果为深入探究蔷薇科植物进化规律奠定了理论基础。
中图分类号:
张冬冬, 韩宏伟, 余镇藩, 曾斌, 杨佳惠, 高雯雯, 马昕彤. 12种蔷薇科植物叶绿体基因组密码子偏好性分析[J]. 中国农业科技导报, 2023, 25(8): 65-75.
Dongdong ZHANG, Hongwei HAN, Zhenfan YU, Bin ZENG, Jiahui YANG, Wenwen GAO, Xintong MA. Codon Preference Analysis of Chloroplast Genome in 12 Rosaceae Plants[J]. Journal of Agricultural Science and Technology, 2023, 25(8): 65-75.
| 属 Genus | 物种 Species | NCBI登录号 Accession number in the NCBI database | 长度 Length/bp |
|---|---|---|---|
| 木瓜属Chaenomeles | 木瓜Chaenomeles sinensis | KT932967.1 | 159 351 |
| 枇杷属Eriobotrya Lindl. | 枇杷Eriobotrya japonica | KT633951.1 | 159 137 |
| 草莓属Fragaria L. | 草莓Fragaria × ananassa | KY358226.1 | 155 549 |
| 苹果属Malus Mill. | 苹果Malus domestica | MK434916.1 | 159 926 |
| 西府海棠Malus micromalus | MF062434.1 | 159 834 | |
| 杏属Armeniaca Mill. | 杏Prunus armeniaca | MH700954.1 | 158 070 |
| 梅花Prunus mume | MH727486.1 | 157 783 | |
| 桃属Amygdalus L. | 扁桃Prunus dulcis | MH700953.1 | 157 916 |
| 桃Prunus persica | HQ336405.1 | 157 790 | |
| 樱属Cerasus Mill. | 樱桃Prunus pseudocerasus | KX255667.1 | 157 834 |
| 李属Prunus L. | 中国李Prunus salicina | MH700952.1 | 157 916 |
| 梨属Pyrus | 西洋梨Pyrus communis | MN577870.1 | 160 171 |
表1 本研究所收集的12种蔷薇科叶绿体基因组
Table 1 Chloroplast genomes of 12 Rosaceae collected in this study
| 属 Genus | 物种 Species | NCBI登录号 Accession number in the NCBI database | 长度 Length/bp |
|---|---|---|---|
| 木瓜属Chaenomeles | 木瓜Chaenomeles sinensis | KT932967.1 | 159 351 |
| 枇杷属Eriobotrya Lindl. | 枇杷Eriobotrya japonica | KT633951.1 | 159 137 |
| 草莓属Fragaria L. | 草莓Fragaria × ananassa | KY358226.1 | 155 549 |
| 苹果属Malus Mill. | 苹果Malus domestica | MK434916.1 | 159 926 |
| 西府海棠Malus micromalus | MF062434.1 | 159 834 | |
| 杏属Armeniaca Mill. | 杏Prunus armeniaca | MH700954.1 | 158 070 |
| 梅花Prunus mume | MH727486.1 | 157 783 | |
| 桃属Amygdalus L. | 扁桃Prunus dulcis | MH700953.1 | 157 916 |
| 桃Prunus persica | HQ336405.1 | 157 790 | |
| 樱属Cerasus Mill. | 樱桃Prunus pseudocerasus | KX255667.1 | 157 834 |
| 李属Prunus L. | 中国李Prunus salicina | MH700952.1 | 157 916 |
| 梨属Pyrus | 西洋梨Pyrus communis | MN577870.1 | 160 171 |
| 物种Specie | CP | T3 | C3 | A3 | G3 | GC1 | GC2 | GC3 | GCall |
|---|---|---|---|---|---|---|---|---|---|
| 木瓜Chaenomeles sinensis | 31.22 | 27.91 | 45.57 | 37.76 | 29.74 | 46.98 | 39.47 | 28.28 | 36.66 |
| 枇杷Eriobotrya japonica | 31.19 | 27.92 | 45.56 | 37.79 | 29.73 | 46.94 | 39.50 | 28.27 | 36.72 |
| 草莓Fragaria × ananassa | 31.66 | 28.57 | 45.60 | 37.73 | 30.62 | 47.06 | 39.45 | 29.29 | 37.23 |
| 苹果Malus domestica | 31.22 | 27.90 | 45.56 | 37.75 | 29.73 | 47.00 | 39.43 | 28.32 | 36.57 |
| 西府海棠Malus micromalus | 31.77 | 28.29 | 45.46 | 37.66 | 29.80 | 46.97 | 39.44 | 28.28 | 36.61 |
| 杏Prunus armeniaca | 31.76 | 28.20 | 45.36 | 37.62 | 29.81 | 46.86 | 39.40 | 28.43 | 36.72 |
| 梅花Prunus mume | 31.26 | 27.86 | 45.48 | 37.76 | 29.89 | 46.90 | 39.40 | 28.61 | 36.76 |
| 扁桃Prunus dulcis | 31.75 | 28.20 | 45.34 | 37.60 | 29.83 | 46.88 | 39.34 | 28.45 | 36.73 |
| 桃Prunus persica | 31.26 | 27.86 | 45.48 | 37.74 | 29.86 | 46.90 | 39.37 | 28.58 | 36.75 |
| 樱桃Prunus pseudocerasus | 31.70 | 28.21 | 45.35 | 37.61 | 29.88 | 46.80 | 39.38 | 28.45 | 36.73 |
| 中国李Prunus salicina | 31.74 | 28.21 | 45.39 | 37.61 | 29.80 | 46.90 | 39.36 | 28.45 | 36.75 |
| 西洋梨Pyrus communis | 31.75 | 28.30 | 45.47 | 37.67 | 29.83 | 46.98 | 39.46 | 28.35 | 36.54 |
表2 12种蔷薇科植物叶绿体基因组密码子的参数特征 (%)
Table 2 Parameter characteristics of codon in chloroplast genome of 12 rosaceae species
| 物种Specie | CP | T3 | C3 | A3 | G3 | GC1 | GC2 | GC3 | GCall |
|---|---|---|---|---|---|---|---|---|---|
| 木瓜Chaenomeles sinensis | 31.22 | 27.91 | 45.57 | 37.76 | 29.74 | 46.98 | 39.47 | 28.28 | 36.66 |
| 枇杷Eriobotrya japonica | 31.19 | 27.92 | 45.56 | 37.79 | 29.73 | 46.94 | 39.50 | 28.27 | 36.72 |
| 草莓Fragaria × ananassa | 31.66 | 28.57 | 45.60 | 37.73 | 30.62 | 47.06 | 39.45 | 29.29 | 37.23 |
| 苹果Malus domestica | 31.22 | 27.90 | 45.56 | 37.75 | 29.73 | 47.00 | 39.43 | 28.32 | 36.57 |
| 西府海棠Malus micromalus | 31.77 | 28.29 | 45.46 | 37.66 | 29.80 | 46.97 | 39.44 | 28.28 | 36.61 |
| 杏Prunus armeniaca | 31.76 | 28.20 | 45.36 | 37.62 | 29.81 | 46.86 | 39.40 | 28.43 | 36.72 |
| 梅花Prunus mume | 31.26 | 27.86 | 45.48 | 37.76 | 29.89 | 46.90 | 39.40 | 28.61 | 36.76 |
| 扁桃Prunus dulcis | 31.75 | 28.20 | 45.34 | 37.60 | 29.83 | 46.88 | 39.34 | 28.45 | 36.73 |
| 桃Prunus persica | 31.26 | 27.86 | 45.48 | 37.74 | 29.86 | 46.90 | 39.37 | 28.58 | 36.75 |
| 樱桃Prunus pseudocerasus | 31.70 | 28.21 | 45.35 | 37.61 | 29.88 | 46.80 | 39.38 | 28.45 | 36.73 |
| 中国李Prunus salicina | 31.74 | 28.21 | 45.39 | 37.61 | 29.80 | 46.90 | 39.36 | 28.45 | 36.75 |
| 西洋梨Pyrus communis | 31.75 | 28.30 | 45.47 | 37.67 | 29.83 | 46.98 | 39.46 | 28.35 | 36.54 |

图2 CN、GC1、GC2、GC3、GCall以及ENC之间相关性注:蓝色表示正相关;橘色表示负相关;颜色越深、圆形越大表示相关性越高。
Fig. 2 Correlation between CN, GC1, GC2, GC3, GCall and ENCNote: Blue indicates positive correlation; orange indicates negative correlation; the dot with darker color and larger circle indicates higher correlation.
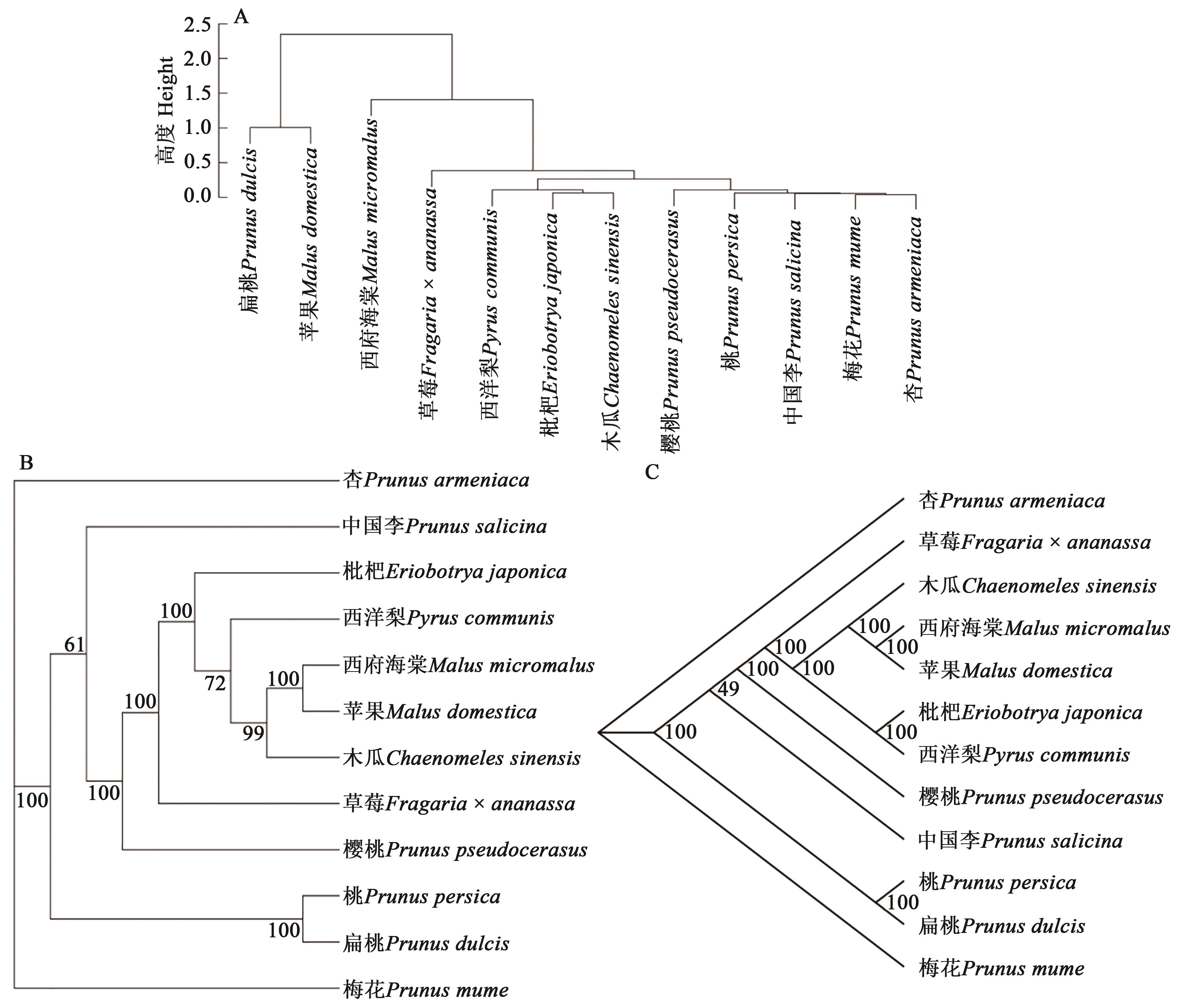
图7 12种蔷薇科植物的系统进化树A:基于RSCU;B:基于53个编码基因序列; C:基于叶绿体基因序列
Fig. 7 Phylogenetic tree of 12 Rosaceae speciesA: Based on RSCU values; B: Based on protein-coding sequences; C: Based on chloroplast gene sequences
| 1 | XU C, DONG W P, LI W Q, et al.. Comparative analysis of six Lagerstroemia complete chloroplast genomes [J/OL]. Front. Plant Sci., 2017, 8:15 [2021-09-10]. . |
| 2 | 吴静妍,葛新新,李海英,等.蔷薇科果树组织培养研究进展[J].安徽农业科学,2020,48(21):10-12. |
| WU J Y, GE X X, LI H Y, et al.. Research progress on tissue culture of Rosaceae fruit trees [J]. J. Anhui Agric. Sci., 2020, 48(21):10-12. | |
| 3 | 刘玉,刘聚栋.月季在园林布景中的应用[J].乡村科技,2021,12(24):106-107. |
| 4 | GUTTERIDGE S. The impact of a changing atmosphere on chloroplast function, photosynthesis, yield, and food security [J]. Essays Biol., 2018, 62(1):1-11. |
| 5 | 陈柯伊,李朝娜,成敏敏,等.不同叶色矢竹叶绿体结构和光系统特性差异[J].植物学报,2018,53(4):509-518. |
| CHEN K Y, LI Z N, CHENG M M, et al.. Chloroplast ultrastructure and chlorophyll fluorescence characteristics of three cultivars of Pseudosasa japonica [J]. Chin. Bull. Bot., 2020, 48(21):10-12. | |
| 6 | NAKAMURA M, SUGIURA M. Translation efficiencies of synonymous codons for arginine differ dramatically and are not correlated with codon usage in chloroplasts [J]. Gene, 2011, 472(1-2):50-54. |
| 7 | DINAKAR C, QINGWEI Z, DOROTHEA B. Protection of photosynthesis in desiccation-tolerant resurrection plants [J]. J. Plant Physiol., 2018, 227:84-92. |
| 8 | ANDREW W G, SHEIL A A, LI Z Z, et al.. Comparative genomics of 11 complete chloroplast genomes of Senecioneae (Asteraceae) species: DNA barcodes and phylogenetics [J/OL]. Bot. Stud., 2019, 60(1):17 [2021-09-10]. . |
| 9 | WEI Q K, JIN H Y. The complete chloroplast genome sequence of Morus cathayana and Morus multicaulis, and comparative analysis within genus Morus L [J/OL]. Peer J., 2017, 5:e3037 [2021-09-10]. . |
| 10 | SUZUKI H, MORTON B R. Codon adaptation of plastid genes [J/OL]. PLoS ONE, 2016, 11(5):e0154306 [2021-09-10]. . |
| 11 | NAKAMURA M, SUGIURA M. Translation efficiencies of synonymous codons for arginine differ dramatically and are not correlated with codon usage in chloroplasts [J]. Gene, 2011, 472(1-2):50-54. |
| 12 | PURABI M, ROFINA Y B, KATHARINA M N, et al.. Codon usage and codon pair patterns in non-grass monocot genomes [J]. Ann. Bot., 2017, 120:893-909. |
| 13 | LIU M Y, DONG H, WANG M, et al.. Evolutionary divergence of function and expression of laccase genes in plants [J/OL]. J. Genet., 2020, 99(1):23 [2021-09-10]. . |
| 14 | LIU X Y, LI Y, JI K K, et al.. Genome-wide codon usage pattern analysis reveals the correlation between codon usage bias and gene expression in Cuscuta australis [J]. Genomics, 2020, 112 (4):2695-2702. |
| 15 | JENSEN P E, LEISTER D. Chloroplast evolution, structure and functions [J/OL]. F1000Prime Rep., 2014, 6:40 [2021-09-10]. . |
| 16 | ZHANG D Z, RABIA B P, LIU J J, et al.. Morphological diversity and correlation analysis of phenotypes and quality traits of proso millet (Panicum miliaceum L.) core collections [J]. J. Integr. Agric., 2019, 18(5):958-969. |
| 17 | HD B, DONG H, JIANG C, et al.. Analysis of codon usage patterns in Ginkgo biloba reveals codon usage tendency from A/U-ending to G/C-ending [J/OL]. Sci. Rep., 2016, 6(1):35927 [2021-09-10]. . |
| 18 | WRIGHT F. The ‘effective number of codons’ used in a gene [J]. Gene, 1990, 87(1):23-29. |
| 19 | SHIVANI G, PATRA P K, YADAV M K. New insights into the factors affecting synonymous codon usage in human infecting Plasmodium species [J]. Acta Trop., 2017, 176:29-33. |
| 20 | RENSING S, FRITZOWSKY D, LANG D, et al.. Protein encoding genes in an ancient plant: analysis of codon usage, retained genes and splice sites in a moss, Physcomitrella patens [J/OL]. BMC Genomics., 2005, 6(1):43 [2021-09-10]. . |
| 21 | 吴宪明,吴松锋,任大明,等.密码子偏性的分析方法及相关研究进展[J].遗传,2007(4):420-426. |
| WU X M, WU S F, REN D M, et al.. The analysis method and progress in the study of codon bias [J]. Hereditas., 2007(4):420-426. | |
| 22 | SONG H, LIU J, SONG Q, et al.. Comprehensive analysis of codon usage bias in seven Epichloë species and their peramine-coding genes [J/OL]. Front. Microbiol., 2017, 8:1419 [2021-09-10]. . |
| 23 | ZHANG Y, NIE X, JIA X, et al.. Analysis of codon usage patterns of the chloroplast genomes in the Poaceae family [J/OL]. Aust. J. Bot., 2012, 60 (5):461 [2021-09-10]. . |
| 24 | MCLEAN M J, WOLFE K H, DEVINE K M. Base composition skews, replication orientation, and gene orientation in 12 prokaryote genomes [J]. J. Mol. Evol., 1998, 47(6):691-696. |
| 25 | 唐玉娟,赵英,黄国弟,等.芒果叶绿体基因组密码子使用偏好性分析[J].热带作物学报,2021,42(8):2143-2150. |
| TANG Y J, ZHAO Y, HUANG G D, et al.. Analysis on codon usage bias of chloroplast genes from mango [J]. Chin. J. Tropical Crop, 2021, 42(8):2143-2150. | |
| 26 | 王鹏良,杨利平,吴红英,等.普通油茶叶绿体基因组密码子偏好性分析[J].广西植物,2018,38(2):135-144. |
| WANG P L, YANG L P, WU H Y, et al.. Condon preference of chloroplast genome in Camellia oleifera [J]. Guihaia, 2018, 38(2):135-144. | |
| 27 | 刘汉梅,赵耀,顾勇,等.几种植物waxy基因的密码子用法特性分析[J].核农学报,2010,24(3):476-481. |
| LIU H M, ZHAO Y, GU Y, et al.. Characterization of codon usage of waxy genes in several plants [J]. J. Nucl. Agric. Sci., 2010, 24(3):476-481. | |
| 28 | KUMAR S, STECHER G, LI M, et al.. MEGA X: molecular evolutionary genetics analysis across computing platforms [J]. Mol. Biol. Evol., 2018, 35(6):1547-1549. |
| 29 | LI S C, CHANG L, ZHANG J. Advancing organelle genome transformation and editing for crop improvement [J/OL]. Plant Commun., 2021, 2(2):100141 [2021-09-10]. . |
| 30 | CHAKRABORTY S, NAG D, MAZUMDER T H, et al.. Codon usage pattern and prediction of gene expression level in Bungarus species [J]. Gene, 2017, 604:48-60. |
| 31 | 朱婷婷,张磊,陈万生,等.1342个植物叶绿体基因组分析[J].基因组学与应用生物学, 2017,36(10):4323-4333. |
| ZHU T T, ZHANG L, CHEN W S, et al.. Analysis of chloroplast genomes in 1342 plants [J]. Genomics Appl. Biol., 2017, 36(10):4323-4333. | |
| 32 | 季凯凯,宋希强,陈春国,等.木兰科叶绿体基因组的密码子使用特征分析[J].中国农业科技导报,2020,22(11):52-62. |
| JI K K, SONG X Q, CHEN C G, et al.. Codon usage profiling of chloroplast genome in Magnoliaceae [J]. J. Agric. Sci. Technol., 2020, 22(11):52-62. | |
| 33 | WANG Z, XU B, LI B, et al.. Comparative analysis of codon usage patterns in chloroplast genomes of six Euphorbiaceae species [J/OL]. Peer J., 2020, 8:e8251 [2021-09-10]. . |
| 34 | LI N, LI Y, ZHENG C, et al.. Genome-wide comparative analysis of the codon usage patterns in plants [J]. Genes Genom., 2016, 38:723-731. |
| 35 | 吴妙丽,陈世品,陈辉.竹亚科叶绿体基因组的密码子使用偏性分析[J].森林与环境学报,2019,39(1):9-14. |
| WU M L, CHEN S P, CHEN H. Condon preference of chloroplast genome of Bambusoideae [J]. J. Fores. Env. 2019, 39(1):9-14. | |
| 36 | 傅建敏,索玉静,刘慧敏,等.柿属植物叶绿体蛋白质编码基因密码子用法[J].经济林研究,2017,35(2):38-44. |
| FU J M, SUO Y J, LIU H M, et al.. Analysis on codon usage in the chloroplast protein-coding genes of Diospyros spp [J]. Nonwood Forest Res., 2017, 35(2):38-44. |
| [1] | 严其伟, 梁湘兰, 陶秋云, 罗秋香, 郭松. 老班瑶药牛耳风叶绿体基因组特征及其密码子使用偏好性分析[J]. 中国农业科技导报, 2022, 24(8): 74-86. |
| [2] | 黄祥, 楚光明, 郑新开, 程锦涛, 陈健豪, 徐迎春, 金奇江, 杨梅花. 睡莲属叶绿体基因组密码子偏好性及系统发育分析[J]. 中国农业科技导报, 2022, 24(4): 75-84. |
| [3] | 苏玥§, 刘娟娟§, 完斌, 张鹏举, 陈正根, 宿俊吉, 王彩香. 乳苣叶绿体基因组特征及其系统发育分析[J]. 中国农业科技导报, 2021, 23(6): 33-42. |
| [4] | 季凯凯1,宋希强1,陈春国2,李革2,谢尚潜1*. 木兰科叶绿体基因组的密码子使用特征分析[J]. 中国农业科技导报, 2020, 22(11): 52-62. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
 京公网安备11010802021197号
京公网安备11010802021197号