




















中国农业科技导报 ›› 2023, Vol. 25 ›› Issue (6): 11-21.DOI: 10.13304/j.nykjdb.2021.0924
收稿日期:2021-11-01
接受日期:2022-01-18
出版日期:2023-06-01
发布日期:2023-07-28
通讯作者:
阎媛媛
作者简介:陈丽婷 E-mail:clt1158433820@163.com;
基金资助:Received:2021-11-01
Accepted:2022-01-18
Online:2023-06-01
Published:2023-07-28
Contact:
Yuanyuan YAN
摘要:
开花是植物产生有性生殖器官的重要阶段,开花整合子整合来自环境和植物体内的信号,进而激活花芽分化,使植物在适宜的时间产生足够多的后代,保障生殖繁衍。棉花中开花整合子基因(FT、SOC1和LFY)已被克隆,但它们在棉花中的进化关系尚不清楚。参考陆地棉基因组,明确了棉属开花整合子同源基因数目,FT和LFY在棉属中进化具有保守性,而SOC1基因序列具有多样性。不同于其他物种,棉花开花整合子基因均在顶端分生组织和叶片中表达量较高,且随着棉苗的生长发育其表达量逐步升高。克隆了GhLFY-2、GhFT-2和GhSOC1-5基因,发现其促进开花的功能是保守的,但开花时间调控机制存在物种特异性。陆地棉中既存在保守的GhFT-2/GhSOC1-5/GhLFY-2线性调控路径,又进化出GhFT-1和GhSOC1-1直接调控GhAP1的路径,表明棉花开花时间调控机制的多样性可能与其对环境的适应性有关。
中图分类号:
陈丽婷, 阎媛媛. 陆地棉开花整合子调控机制解析[J]. 中国农业科技导报, 2023, 25(6): 11-21.
Liting CHEN, Yuanyuan YAN. Investigation of Regulatory Mechanism of Floral Integrators in Upland Cotton[J]. Journal of Agricultural Science and Technology, 2023, 25(6): 11-21.
引物名称 Primer name | 引物序列 Primer sequence(5’-3’) | 用途 Purpose |
|---|---|---|
| GhLFY-2-F(PstI) | TT | 基因克隆 Gene cloning |
| GhLFY-2-R(SmaI) | TT | |
| GhSOC1-4-F(EcoRI) | CC | |
| GhSOC1.4-R(BamHI) | TT | |
| GhFT-2-F(EcoRI) | TT | |
| GhFT-2-R(XmaI) | TT | |
| GhLFY-2-RT-F | AATTCGAGGGTGGTGATGAT | qRT-PCR |
| GhLFY-2-RT-R | CTCCGTTACGATGAAAGGGT | |
| GhSOC1-5-RT-F | GGCGCATAGAGAACGATACA | |
| GhSOC1-5-RT-R | TTTGTATGCCGCCTATAACG | |
| GhFT-2-RT-F | GGGATGATCTGAGGACCTTCT | |
| GhFT-2-RT-R | CACCAGTTGTGGCTGGAATA | |
| AtTUB2-F | ATCCGTGAAGAGTACCCAGAT | |
| AtTUB2-R | AAGAACCATGCACTCATCAGC |
表1 本研究中所用引物序列
Table 1 Primer sequences in the study
引物名称 Primer name | 引物序列 Primer sequence(5’-3’) | 用途 Purpose |
|---|---|---|
| GhLFY-2-F(PstI) | TT | 基因克隆 Gene cloning |
| GhLFY-2-R(SmaI) | TT | |
| GhSOC1-4-F(EcoRI) | CC | |
| GhSOC1.4-R(BamHI) | TT | |
| GhFT-2-F(EcoRI) | TT | |
| GhFT-2-R(XmaI) | TT | |
| GhLFY-2-RT-F | AATTCGAGGGTGGTGATGAT | qRT-PCR |
| GhLFY-2-RT-R | CTCCGTTACGATGAAAGGGT | |
| GhSOC1-5-RT-F | GGCGCATAGAGAACGATACA | |
| GhSOC1-5-RT-R | TTTGTATGCCGCCTATAACG | |
| GhFT-2-RT-F | GGGATGATCTGAGGACCTTCT | |
| GhFT-2-RT-R | CACCAGTTGTGGCTGGAATA | |
| AtTUB2-F | ATCCGTGAAGAGTACCCAGAT | |
| AtTUB2-R | AAGAACCATGCACTCATCAGC |

图2 植物开花整合子基因的进化树注:Gh—陆地棉;Gb—海岛棉;Ga—亚洲棉;Gr—雷蒙德氏棉;At—拟南芥;Pt—杨树;Nt—烟草;Vv—葡萄。
Fig. 2 Phylogenetic analysis of plant floral integrator genesNote:Gh—Gossypium hirsutum; Gb—Gossypium barbadense; Ga—Gossypium arboreum; Gr—Gossypium raimondii; At—Arabidopsis thaliana; Pt—Populus tomentosa; Nt—Nicotiana tabacum; Vv—Vitis vinifera.
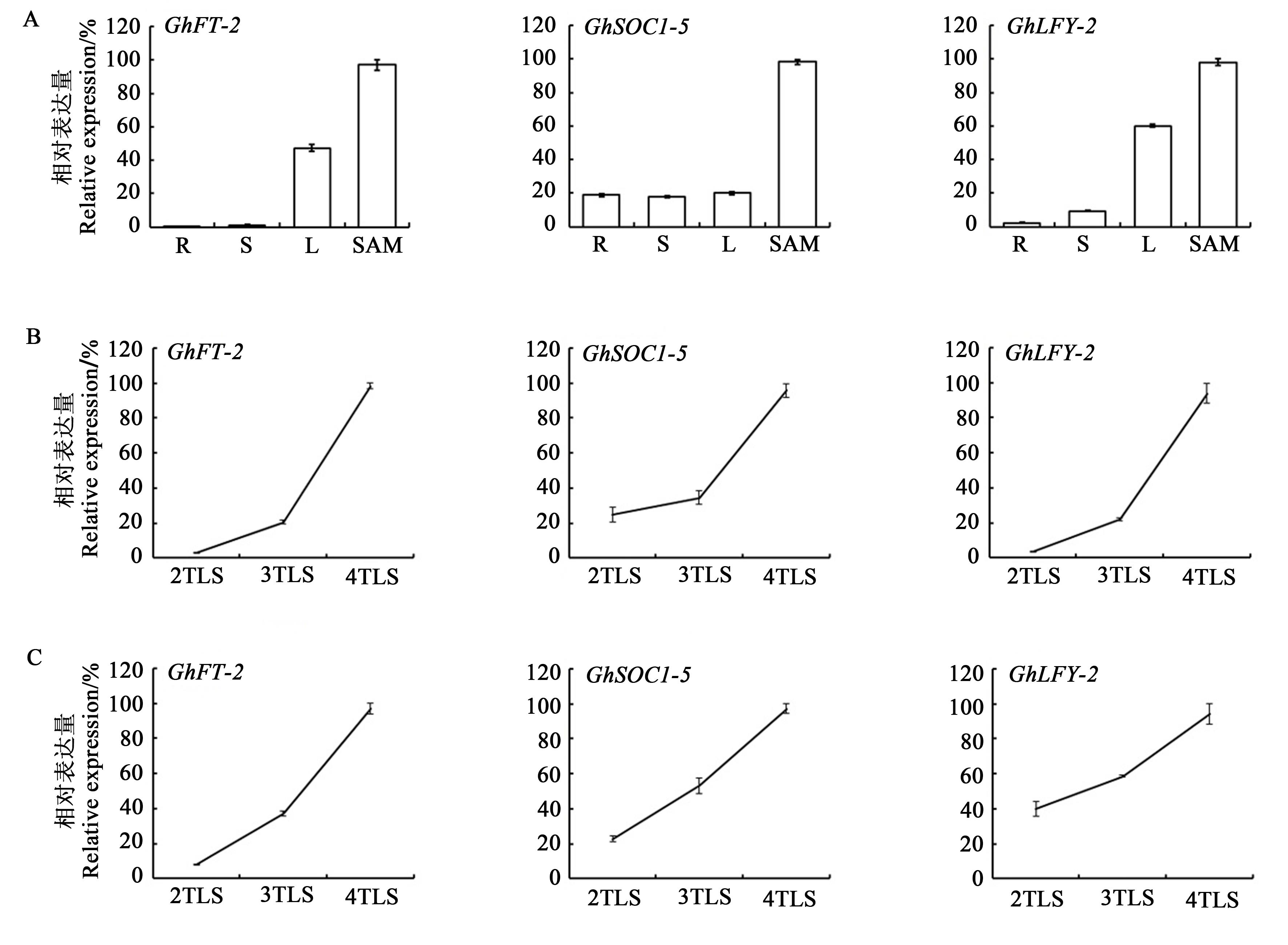
图3 开花整合子基因在陆地棉中表达A:棉花开花整合子基因组织特异性表达;B:棉花开花整合子基因随棉苗生长在叶片中表达变化;C:棉花开花整合子基因随棉苗生长在SAM中表达变化。R—根;S—茎;L—叶;TLS—真叶期
Fig. 3 Expression of floral integrators genes in Gossypium hirsutumA: Tissue-specific expression of flowering integrator of Gossypium hirsutum; B: Expression trends of flowering integrators in cotton leaves during seedling development; C: Expression trends of flowering integrators in cotton SAM during seedling development. R—Root; S—Stem; L—Leaf; TLS—True leaf stage

图4 棉花开花整合子促进开花A:转基因拟南芥早花表型;B:转基因拟南芥基因的表达量及莲座叶数目统计;C:野生型和转基因拟南芥的开花时间,**表示与野生型相比差异显著(P<0.01);D:GhLFY-2转基因植株莲座叶叶腋里伸出单生花
Fig. 4 Cotton flowering integrators promotes floweringA: Early flowering phenotype of transgenic plants; B: Comparison of gene expression and flowering time between WT and transgenic plants; C: Flowering time of WT and transgenic lines, ** indicates significant differences (P<0.01); D: Solitary flowers produced in GhLFY-2 transgenic plants

图5 拟南芥开花整合子基因转录水平注:A、B和C分别为开花整合子在GhFT-2、GhSOC1-5和GhLFY-2转基因株系中的表达量。
Fig. 5 Expression of floral integrator genes in transgenic Arabidopsis plantsNote:A, B and C shows relative expression of flowering integrators in GhFT-2, GhSOC1-5 and GhLFY-2 transgenic plants, respectively.
| 1 | 喻树迅,王寒涛,魏恒玲,等.棉花早熟性研究进展及其应用[J].棉花学报, 2017, 29(): 1-10. |
| YU S X, WANG H T, WEI H L, et al.. Research progress and application of early maturity in upland cotton [J]. Cotton Sci., 2017, 29(S1): 1-10. | |
| 2 | FORNARA F, MONTAIGU A D, COUPLAND G. SnapShot: control of flowering in Arabidopsis [J]. Cell, 2010, 141(3): 550-550. |
| 3 | WIGGE P. FT, a mobile developmental signal in plants [J]. Curr. Biol., 2011, 21(9): R374-R378. |
| 4 | KOBAYASHI Y, KAYA H, GOTO K, et al.. A pair of related genes with antagonistic roles in mediating flowering signals [J]. Science, 2000, 286(5446): 1960-1962. |
| 5 | ABE M, KOBAYASHI Y, YAMAMOTO S, et al.. FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex [J]. Science, 2005, 309(5737): 1052-1056. |
| 6 | WIGGE P A, MIN C K, JAEGER K E, et al.. Integration of spatial and temporal information during floral induction in Arabidopsis [J]. Science, 2005, 309(5737): 1056-1059. |
| 7 | TAMAKI S, MATSUO S, WONG H L, et al.. Hd3a protein is a mobile flowering signal in rice [J]. Science, 2007, 316(5827): 1033-1036. |
| 8 | MCGARRY R C, PREWITT S, AYRE B G. Overexpression of FT in cotton affects architecture but not floral organogenesis [J/OL]. Plant Signal Behav., 2013, 8(4):e23602 [2021-12-03]. . |
| 9 | GUO D, LI C, DONG R, et al.. Molecular cloning and functional analysis of the FLOWERING LOCUS T(FT) homolog GhFT1 from Gossypium hirsutum [J]. J. Integr. Plant Biol., 2015, 57(6): 522-533. |
| 10 | LI C, ZHANG Y, ZHANG K, et al.. Promoting flowering, lateral shoot outgrowth, leaf development, and flower abscission in tobacco plants overexpressing cotton FLOWERING LOCUS T (FT)-like gene GhFT1 [J/OL]. Front. Plant Sci., 2015, 6: 454 [2021-12-03]. . |
| 11 | BLÁZQUEZ M A, WEIGEL D. Integration of floral inductive signals in Arabidopsis [J]. Nature, 2000, 404(6780): 889-892. |
| 12 | YOO S K, CHUNG K S, KIM J, et al.. Constant activates suppressor of overexpression of constant 1 through flowering locus T to promote flowering in Arabidopsis [J]. Plant Physiol., 2005, 139(2): 770-778. |
| 13 | ZHANG X, WEI J, FAN S, et al.. Functional characterization of GhSOC1 and GhMADS42 homologs from upland cotton (Gossypium hirsutum L.) [J]. Plant Sci., 2016, 242: 178-186. |
| 14 | CERDÁN P D, CHORY J. Regulation of flowering time by light quality [J]. Nature, 2003, 423(6942):881-885. |
| 15 | SVEN E, HENRIK B, THOMAS M, et al.. GA4 is the active gibberellin in the regulation of LEAFY transcription and Arabidopsis floral initiation [J]. Plant Cell, 2006, 18(9): 2172-2181. |
| 16 | LILJEGREN S J, GUSTAFSON-BROWN C, PINYOPICH A, et al.. Interactions among APETALA1, LEAFY, and TERMINAL FLOWER1 specify meristem fate [J]. Plant Cell, 1999, 11(6): 1007-1018. |
| 17 | WAGNER D, SABLOWSKI R, MEYEROWITZT E M. Transcriptional activation of APETALA1 by LEAFY [J]. Science, 1999, 285(5427): 582-584. |
| 18 | HEMPEL F D, WEIGEL D, MANDEL M A, et al.. Floral determination and expression of floral regulatory genes in Arabidopsis [J]. Development, 1997, 124(19): 3845-3853. |
| 19 | BLAZQUEZ M A, SOOWAL L N, LEE I, et al.. LEAFY expression and flower initiation in Arabidopsis [J]. Development, 1997, 124(19): 3835-3844. |
| 20 | WEIGEL D, NILSSON O. A developmental switch sufficient for flower initiation in diverse plants [J]. Nature, 1995, 377(6549): 495-500. |
| 21 | AHEARN K P, JOHNSON H A, DETLEF W, et al.. NFL1, a Nicotiana tabacum LEAFY-like gene, controls meristem initiation and floral structure [J]. Plant Cell Physiol., 2001, 10: 1130-1139. |
| 22 | KYOZUKA J, KONISHI S, NEMOTO K, et al.. Down-regulation of RFL, the FLO/LFY homolog of rice, accompanied with panicle branch initiation [J]. Proc. Natl. Acad. Sci. USA, 1998, 95(5): 1979-1982. |
| 23 | LI J, FAN S L, SONG M Z, et al.. Cloning and characterization of a FLO/LFY ortholog in Gossypium hirsutum L [J]. Plant Cell Rep., 2013, 32(11): 1675-1686. |
| 24 | LEE J, OH M, PARK H, et al.. SOC1 translocated to the nucleus by interaction with AGL24 directly regulates LEAFY [J]. Plant J., 2008, 55(5): 832-843. |
| 25 | YAN Y, SHEN L, CHEN Y, et al.. A MYB-domain protein EFM mediates flowering responses to environmental cues in Arabidopsis [J]. Dev. Cell, 2014, 30(4): 437-448. |
| 26 | HIGUCHI Y, NARUMI T, ODA A, et al.. The gated induction system of a systemic floral inhibitor, antiflorigen, determines obligate short-day flowering in chrysanthemums [J]. Proc. Natl. Acad. Sci. USA, 2013, 110(42): 17137-17142. |
| 27 | SONG Q, ZHANG T, STELLY D M, et al.. Epigenomic and functional analyses reveal roles of epialleles in the loss of photoperiod sensitivity during domestication of allotetraploid cottons [J/OL]. Genome Biol., 2017, 18(1): 99 [2021-12-03]. . |
| 28 | MA Z, ZHANG Y, WU L . et al ..High-quality genome assembly and resequencing of modern cotton cultivars provide resources for crop improvement [J]. Nat. Genet., 2021, 53: 1385-1391. |
| 29 | LIVAK K J, SCHMITTGEN T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method [J]. Methods, 2001, 25(4): 402-408. |
| 30 | SARAH M N, SINARA A, GUSTAVO M A, et al.. Genome-wide analysis of the MADS-box gene family in polyploid cotton (Gossypium hirsutum) and in its diploid parental species (Gossypium arboreum and Gossypium raimondii) [J]. Plant Physiol. Bioch., 2018, 127: 169-184. |
| 31 | REN Z Y, YU D Q, YANG Z E, et al.. Genome-wide identification of the MIKC-Type MADS-box gene family in Gossypium hirsutum L. unravels their roles in flowering [J/OL]. Front. Plant Sci., 2017, 8: 384 [2021-12-03]. . |
| 32 | PATIRANAGE D, ASARE E, MALDONADO-TAIPE N, et al.. Haplotype variations of major flowering time genes in quinoa unveil their role in the adaptation to different environmental conditions [J]. Plant Cell Environ., 2021, 44(8): 2565-2579. |
| 33 | 樊红娟,程雪敏,郑思,等.棉花GhFT基因的克隆及功能分析[J].分子植物育种, 2019, 18(17): 29-36. |
| FAN H J, CHENG X M, ZHENG S, et al.. The cloning and function analysis of GhFT gene in Gossypium hirsutum [J]. Mol. Plant Breed., 2020, 18(17): 29-36. | |
| 34 | JACK T. Molecular and genetic mechanisms of floral control [J]. Plant Cell, 2004, 16 (Supp. l): S1-S17. |
| 35 | MAIZEL A, BUSCH M A, TANAHASHI T, et al.. The floral regulator LEAFY evolves by substitutions in the DNA binding domain [J]. Science, 2005, 308(5719): 260-263. |
| 36 | KELLY A J, BONNLANDER M B, MEEKS-WAGNER D R. NFL, the tobacco homolog of FLORICAULA and LEAFY, is transcriptionally expressed in both vegetative and floral meristems [J]. Plant Cell, 1995, 7(2): 225-234. |
| 37 | HOFER J, TURNER L, HELLENS R, et al.. UNIFOLIATA regulates leaf and flower morphogenesis in pea [J]. Curr. Biol., 1997, 7(8): 581-587. |
| 38 | CARMONA M J, CUBAS P, MARTÍNEZ-ZAPATER J M. VFL, the grapevine FLORICAULA/LEAFY ortholog, is expressed in meristematic regions independently of their fate [J]. Plant Physiol., 2002, 130(1): 68-77. |
| [1] | 张曼, 王志城, 刘正文, 王国宁, 王省芬, 张艳. 陆地棉BGLU基因家族成员的全基因组鉴定与表达分析[J]. 中国农业科技导报, 2023, 25(2): 48-59. |
| [2] | 李美丽, 宿俊吉, 杨永林, 秦江鸿, 李鲜鲜, 杨德龙, 马麒, 王彩香. 陆地棉COI家族基因鉴定及在干旱和盐胁迫下的表达分析[J]. 中国农业科技导报, 2022, 24(4): 63-74. |
| [3] | 默韶京, 王志城, 王省芬, 刘正文, 吴立强, 张桂寅, 马峙英, 张艳, 段会军. 陆地棉GELP家族基因鉴定及其响应胁迫的表达分析[J]. 中国农业科技导报, 2022, 24(2): 93-103. |
| [4] | 郑巨云, 桑志伟, 王俊铎, 龚照龙, 梁亚军, 张泽良, 郭江平, 莫明, 李雪源. 棉花品种抗旱性相关指标分析与综合评价[J]. 中国农业科技导报, 2022, 24(10): 23-34. |
| [5] | 王琴琴, 陈修贵, 陆许可, 王帅, 张悦新, 范亚朋, 陈全家, 叶武威. 陆地棉GhPKE1的生物信息学分析及功能验证[J]. 中国农业科技导报, 2022, 24(1): 38-45. |
| [6] | 李生梅, 杨涛, 黄雅婕, 任丹, 耿世伟, 李典鹏, 芮存, 高文伟. 海陆回交群体主要农艺性状与纤维品质关系的探讨[J]. 中国农业科技导报, 2021, 23(8): 16-24. |
| [7] | 张全艳, 张培高, 徐春霞, 丛一宁, 刘丽. 玉米叶夹角的遗传与分子调控研究进展[J]. 中国农业科技导报, 2021, 23(10): 15-24. |
| [8] | 陈斌, 石荣康, 王志城, 刘松, 李青, 刘正文, 孙正文, 王国宁, 吴金华, 马峙英, 张艳, 王省芬. 陆地棉核心种质抗黄萎病鉴定与优异种质筛选[J]. 中国农业科技导报, 2021, 23(10): 45-51. |
| [9] | 王俊铎1,曾辉2,龚照龙1,梁亚军1,艾先涛1,郭江平1,莫明1,李雪源1,郑巨云1*. 陆地棉品种资源耐复合盐碱性综合评价分析[J]. 中国农业科技导报, 2019, 21(10): 1-11. |
| [10] | 杨笑敏,芮存,张悦新,王德龙,王俊娟,陆许可,陈修贵,郭丽雪,王帅,陈超,叶武威*. 棉花DNA甲基转移酶GhDMT9抗逆性分析[J]. 中国农业科技导报, 2019, 21(10): 12-19. |
| [11] | 王昭玉1,吕婷婷1,李爱芹2,王頔1,张子玉1,闫瑞娴1,. 陆地棉钾吸收相关基因GhHAK1的克隆与功能分析[J]. 中国农业科技导报, 2017, 19(10): 21-27. |
| [12] | 高丽华1,刘博欣1,2,李金博3,吴燕民1,唐益雄1*. 陆地棉bHLH转录因子GhMYC4基因的克隆及功能分析[J]. 中国农业科技导报, 2016, 18(5): 33-41. |
| [13] | 路晓,孙红梅,张伟,李春义*. 以鹿茸为模型探索软骨组织发生[J]. , 2015, 17(4): 71-77. |
| [14] | 孙娜1,王劲2,左开井1*. 陆地棉GhbHLH4基因的克隆与功能分析[J]. , 2014, 16(3): 29-35. |
| [15] | 王莉萍1,孙国清2,梁亚军3,陈全家1*. 棉花纤维品质性状与SSR标记的关联分析[J]. , 2013, 15(4): 110-120. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
 京公网安备11010802021197号
京公网安备11010802021197号 