




















Journal of Agricultural Science and Technology ›› 2023, Vol. 25 ›› Issue (7): 77-86.DOI: 10.13304/j.nykjdb.2022.0170
• BIOTECHNOLOGY & LIFE SCIENCE • Previous Articles Next Articles
Yunsheng WANG( ), Yincui CHEN, Zai CHENG, Jin ZHANG, Chuanbo ZHANG(
), Yincui CHEN, Zai CHENG, Jin ZHANG, Chuanbo ZHANG( )
)
Received:2022-03-07
Accepted:2022-04-20
Online:2023-07-15
Published:2023-08-25
Contact:
Chuanbo ZHANG
通讯作者:
张传博
作者简介:王云胜 E-mail:1710818668@qq.com;
基金资助:CLC Number:
Yunsheng WANG, Yincui CHEN, Zai CHENG, Jin ZHANG, Chuanbo ZHANG. Effects of Overexpression of veA Gene on Secondary Metabolism of Eurotium cristatus[J]. Journal of Agricultural Science and Technology, 2023, 25(7): 77-86.
王云胜, 陈银翠, 程在, 张锦, 张传博. 过表达veA基因对冠突散囊菌次级代谢的影响[J]. 中国农业科技导报, 2023, 25(7): 77-86.
引物 Primer | 序列 Sequence (5’-3’) |
|---|---|
| veAF | AGACATCACCATGGTAGATCTATGGCGACGCGAATGCCC |
| veAR | GGGGAAATTCGAGCTGGTCACCTTAACTAGAAAAAGCCGGCGG |
| GAPDHF | CTGCCGTATCGAGAAGGGTG |
| GAPDHR | GATGAAGTTGGGGTTGAGGG |
| QveAF | CGACATCACCTTTGCCTAC |
| QveAR | TCGGGAAAGATGAAGTAGC |
Table 1 Primers used in this study
引物 Primer | 序列 Sequence (5’-3’) |
|---|---|
| veAF | AGACATCACCATGGTAGATCTATGGCGACGCGAATGCCC |
| veAR | GGGGAAATTCGAGCTGGTCACCTTAACTAGAAAAAGCCGGCGG |
| GAPDHF | CTGCCGTATCGAGAAGGGTG |
| GAPDHR | GATGAAGTTGGGGTTGAGGG |
| QveAF | CGACATCACCTTTGCCTAC |
| QveAR | TCGGGAAAGATGAAGTAGC |
样品 Sample | 原始数据 Raw reads | 质控数据 Clean reads | 错配率 Error rate/% | Q20/% | Q30/% |
|---|---|---|---|---|---|
| OE::veA1_1 | 53 215 236 | 52 880 844 | 0.023 9 | 98.44 | 95.28 |
| OE::veA1_2 | 58 759 294 | 58 355 148 | 0.024 0 | 98.4 | 95.21 |
| OE::veA1_3 | 56 979 242 | 56 535 564 | 0.024 1 | 98.36 | 95.11 |
| WT1 | 52 014 002 | 51 659 080 | 0.024 0 | 98.41 | 95.24 |
| WT2 | 57 594 222 | 57 246 210 | 0.023 9 | 98.43 | 95.26 |
| WT3 | 58 672 768 | 58 282 756 | 0.023 9 | 98.42 | 95.25 |
Table 2 Transcriptome sequencing data statistics
样品 Sample | 原始数据 Raw reads | 质控数据 Clean reads | 错配率 Error rate/% | Q20/% | Q30/% |
|---|---|---|---|---|---|
| OE::veA1_1 | 53 215 236 | 52 880 844 | 0.023 9 | 98.44 | 95.28 |
| OE::veA1_2 | 58 759 294 | 58 355 148 | 0.024 0 | 98.4 | 95.21 |
| OE::veA1_3 | 56 979 242 | 56 535 564 | 0.024 1 | 98.36 | 95.11 |
| WT1 | 52 014 002 | 51 659 080 | 0.024 0 | 98.41 | 95.24 |
| WT2 | 57 594 222 | 57 246 210 | 0.023 9 | 98.43 | 95.26 |
| WT3 | 58 672 768 | 58 282 756 | 0.023 9 | 98.42 | 95.25 |
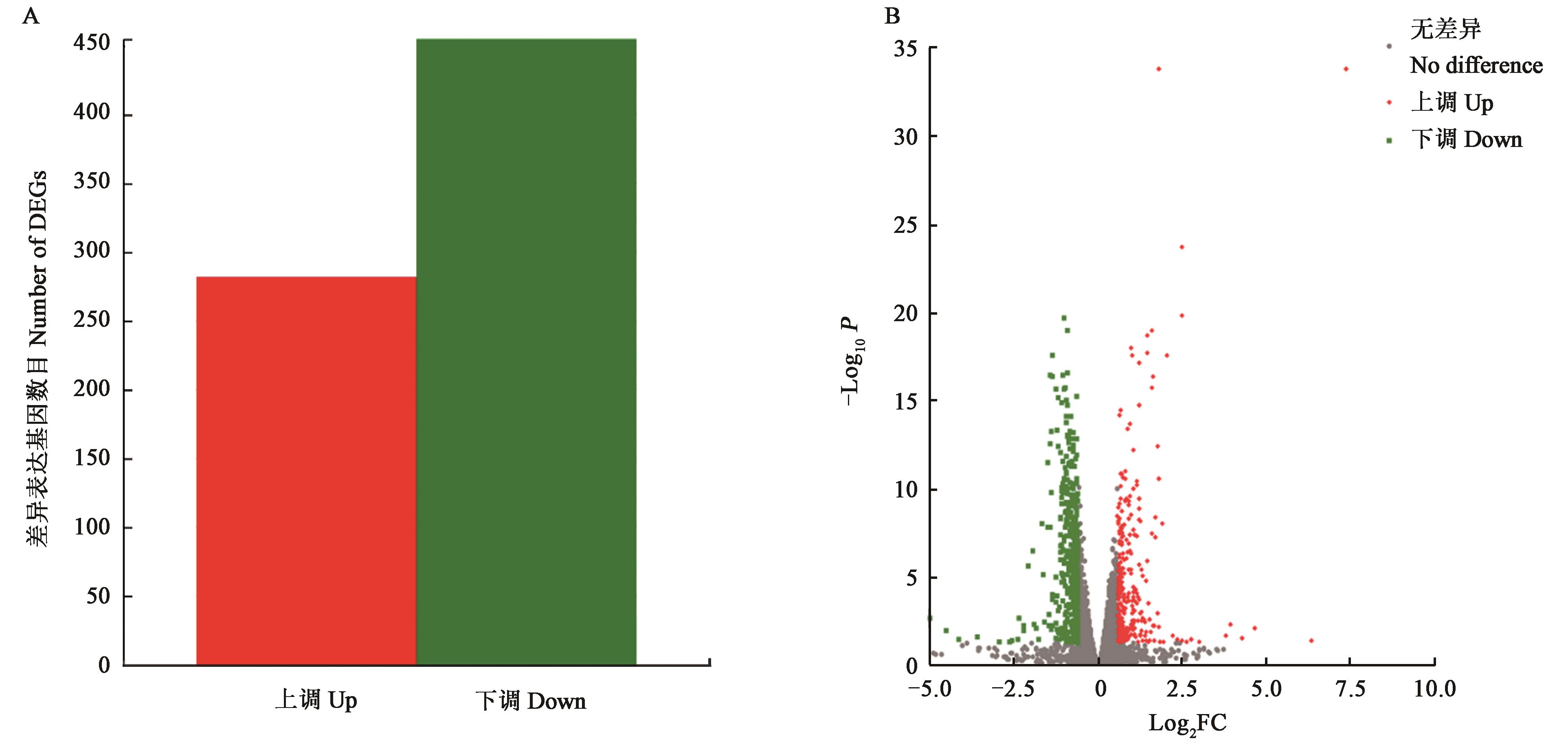
Fig. 3 Differentially expressed genes between WT and OE::veA1A: Histogram of differentially expressed genes; B: Volcano map of differentially expressed genes
基因编号 Gene ID | 功能注释 Function annotation | 变化倍数 Fold change | 调控 Regulate |
|---|---|---|---|
| SI65_05759 | 氧化还原酶 Flavoprotein-ubiquinone oxidoreductase | 1.01ns | 上调Up |
| SI65_05760 | 反转录酶 Reverse transcriptase | 0.61ns | 下调Down |
| SI65_05761 | TATA框结合蛋白 TATA-box-binding protein | 0.72ns | 下调Down |
| SI65_05762 | L-甲基哌啶氧化酶 L-pipecolate oxidase | 1.50* | 上调Up |
| SI65_05763 | 甲基转移酶 Methyltransferase | 1.05ns | 上调Up |
| SI65_05764 | 假设蛋白 Hypothetical protein | 1.45ns | 上调Up |
| SI65_05765 | C6指域转录因子 C6 finger domain transcription factor | 1.25ns | 上调Up |
| SI65_05766 | NADH-生化色素b5还原酶 NADH-cytochrome b5 reductase | 1.27ns | 上调 Up |
| SI65_05767 | 甲基转移酶 Methyltransferase | 1.72* | 上调Up |
| SI65_05768 | 羧基酸合成酶 Carboxylic acid synthase | 2.02* | 上调Up |
| SI65_05769 | 金属化合物-β-内酰胺酶 Metallo-beta-lactamase | 2.31* | 上调Up |
| SI65_05770 | 细胞色素P450单加氧酶 Cytochrome P450 monooxygenase | 1.92ns | 上调Up |
| SI65_05771 | 细胞色素P450单加氧酶 Cytochrome P450 monooxygenase | 2.47* | 上调 Up |
| SI65_05772 | 氧化还原酶 Oxidoreductase | 2.67* | Up 上调 |
| SI65_05773 | 乙酰基转移酶 Acetyltransferase | 2.74* | Up 上调 |
| SI65_05774 | 假定蛋白 Hypothetical protein | NA | NA |
| SI65_05775 | 假定蛋白 Hypothetical protein | 0.55* | Down 下调 |
| SI65_05776 | 转录因子 Transcription factor | 0.64* | Down 下调 |
Table 3 Gene expression and functional annotation in secondary metabolite gene cluster 32
基因编号 Gene ID | 功能注释 Function annotation | 变化倍数 Fold change | 调控 Regulate |
|---|---|---|---|
| SI65_05759 | 氧化还原酶 Flavoprotein-ubiquinone oxidoreductase | 1.01ns | 上调Up |
| SI65_05760 | 反转录酶 Reverse transcriptase | 0.61ns | 下调Down |
| SI65_05761 | TATA框结合蛋白 TATA-box-binding protein | 0.72ns | 下调Down |
| SI65_05762 | L-甲基哌啶氧化酶 L-pipecolate oxidase | 1.50* | 上调Up |
| SI65_05763 | 甲基转移酶 Methyltransferase | 1.05ns | 上调Up |
| SI65_05764 | 假设蛋白 Hypothetical protein | 1.45ns | 上调Up |
| SI65_05765 | C6指域转录因子 C6 finger domain transcription factor | 1.25ns | 上调Up |
| SI65_05766 | NADH-生化色素b5还原酶 NADH-cytochrome b5 reductase | 1.27ns | 上调 Up |
| SI65_05767 | 甲基转移酶 Methyltransferase | 1.72* | 上调Up |
| SI65_05768 | 羧基酸合成酶 Carboxylic acid synthase | 2.02* | 上调Up |
| SI65_05769 | 金属化合物-β-内酰胺酶 Metallo-beta-lactamase | 2.31* | 上调Up |
| SI65_05770 | 细胞色素P450单加氧酶 Cytochrome P450 monooxygenase | 1.92ns | 上调Up |
| SI65_05771 | 细胞色素P450单加氧酶 Cytochrome P450 monooxygenase | 2.47* | 上调 Up |
| SI65_05772 | 氧化还原酶 Oxidoreductase | 2.67* | Up 上调 |
| SI65_05773 | 乙酰基转移酶 Acetyltransferase | 2.74* | Up 上调 |
| SI65_05774 | 假定蛋白 Hypothetical protein | NA | NA |
| SI65_05775 | 假定蛋白 Hypothetical protein | 0.55* | Down 下调 |
| SI65_05776 | 转录因子 Transcription factor | 0.64* | Down 下调 |
| 1 | BOK J, HOFFMEISTER D, MAGGIO-HALL L, et al.. Genomic mining for Aspergillus natural products [J]. Chem. Biol., 2006, 13(1):7-31. |
| 2 | RAJA H A, MILLER A N, PERACE C J, et al.. Fungal identification using molecular tools: a primer for the natural products research community [J]. J. Nat. Prod., 2017, 80(3):756-770. |
| 3 | ZHANG X J, ZHU Y Y, BAO L F, et al.. Putative methyltransferase laea and transcription factor CREA are necessary for proper asexual development and controlling secondary metabolic gene cluster expression [J]. Fungal. Genet. Biol., 2016, 94:32-46. |
| 4 | BAYRAM O, BRAUS G H. Coordination of secondary metabolism and development in fungi: the velvet family of regulatory proteins [J]. FEMS Microbiol. Lett., 2012, 36(1):1-24. |
| 5 | KELLER N. Fungal secondary metabolism: regulation, function and drug discovery [J]. Nat. Rev. Microbiol., 2019, 17(3):167-180. |
| 6 | CHIANG Y M, OAKLEY C E, AHUJA M E, et al.. An efficient system for heterologous expression of secondary metabolite genes in Aspergillus Nidulans [J]. J. Am. Chem. Soc., 2013, 135(20):7720-7731. |
| 7 | YAEGASHI J, OAKLEY B, WANG C. Recent advances in genome mining of secondary metabolite biosynthetic gene clusters and the development of heterologous expression systems in Aspergillus nidulans [J]. J. Appl. Microbiol., 2014, 41(2):433-442. |
| 8 | SARIKAYA-BATRAM Ö, PALMER JM, KELLER N, et al.. One juliet and four romeos: vea and its methyltransferases [J/OL]. Front. Microbiol., 2015, 6:1 [2022-02-10]. . |
| 9 | KATO N, BROOKS W, CALVO A. The expression of sterigmatocystin and penicillin genes in Aspergillus Nidulans is controlled by vea, a gene required for sexual development [J]. Eukaryot. Cell, 2003, 2(6):1178-1186. |
| 10 | MARUI J, OHASHI-KUNIHIRO S, ANDO T, et al.. Penicillin biosynthesis in Aspergillus Oryzae and its overproduction by genetic engineering [J]. J. Biosci. Bioeng., 2010, 110(1):8-11. |
| 11 | 曹双,郑跃亮,张虎,等.过表达veA对橘青霉美伐他汀生物合成的影响[J].西南师范大学学报(自然科学版),2016,41(6):67-73. |
| CAO S, ZHENG Y L, ZHANG H, et al.. On investigation of veA gene in regulating mevastatin biosynthesis, conidia development in Penicillium citrinum [J]. J. Southwest China Normal Univ. (Nat. Sci.), 2016, 41(6):67-73. | |
| 12 | 张月,崔旋旋,刘英学,等.茯砖茶中冠突散囊菌的分离鉴定及其发酵工艺和生物活性研究[J].食品与发酵工业,2020,418(22):202-207. |
| ZHANG Y, CUI X X, LIU Y X, et al.. Isolation, identification, fermentation technology and bioactivity of Eurotium cristatum in Fuzhuan brick tea [J]. Food. Ferment. Ind., 2020, 46(22):202-207. | |
| 13 | AN T T, CHEN M X, ZU Z Q, et al.. Untargeted and targeted metabolomics reveal changes in the chemical constituents of instant dark tea during liquid-state fermentation by Eurotium cristatum [J]. Food. Res. Int., 2021, 148:110-125. |
| 14 | 陈敏,谢发,游玲,等.冠突散囊菌发酵苦丁茶工艺研究[J].食品与发酵工业, 2020,402(6):224-228. |
| CHNEG M, XIE F, YOU L, et al.. Fermentation of ligustrum robustum by Eurotium cristatum [J]. Food. Ferment. Ind., 2020, 46(6):224-228. | |
| 15 | GE Y Y, WANG Y C, LIU Y X, et al.. Comparative genomic and transcriptomic analyses of the Fuzhuan brick tea-fermentation fungus Aspergillus cristatus [J/OL]. BMC Genomics, 2016, 17(1): 428 [2022-02-10]. . |
| 16 | 周国庆.冠突曲霉LaeA基因功能的研究[D].贵阳:贵州大学,2018. |
| ZHOU G Q. Study on the function of LaeA gene in Aspergillus cristatum [D]. Guiyang: Guizhou University, 2018. | |
| 17 | TAN Y M, WANG H, WANG Y P, et al.. The role of the vea gene in adjusting developmental balance and environmental stress response in Aspergillus cristatus [J]. Fungal. Biol., 2018, 122(10):952-964. |
| 18 | WANG Y P, TAN Y M, WANG Y C, et al.. Role of Acndta in cleistothecium formation, osmotic stress response, pigmentation and carbon metabolism of Aspergillus cristatus [J]. Fungal. Biol., 2021, 125(10):749-763. |
| 19 | 刘逸梅.冠突曲霉钙信号调控基因Fig的功能研究[D].贵阳:贵州大学, 2019. |
| LIU YM. Study on the function of Fig gene regulating calcium signal in Aspergillus cristatum [D]. Guiyang: Guizhou University, 2019. | |
| 20 | 朱思远,徐岩,喻晓蔚.农杆菌介导转化黑曲霉条件优化及脂肪酶表达[J].食品与生物技术学报,2020,39(5):51-58. |
| ZHU S Y, XU Y, YU X Y. Optimization of agrobacterium tumefaciens-mediated transformation of Aspergillus niger and expression of lipase [J]. J. Food Sci. Biotechnol., 2020, 39(5):51-58. | |
| 21 | 邵蕾,谭玉梅,任春光,等.转录因子ACZ影响冠突曲霉色素及产孢数量的研究[J].基因组学与应用生物学,2019,38(5):2041-2048. |
| SHAO L, TAN Y M, REN C G, et al.. Study on the transcription factor ACZ affecting the number of spores and pigmentation of Asperillus cristatus [J]. Genom. Appl. Biol., 2019, 38(5):2041-2048. | |
| 22 | 向婷,于凤明,杨正军,等.冠突曲霉preA基因的功能分析[J].基因组学与应用生物学,2020,39(7):3070-3078. |
| XIANG T, YU F M, YANG Z J, et al.. Functional analysis on the preA gene in Aspergillus cristatus [J]. Genom. Appl. Biol., 2020, 39(7):3070-3078. | |
| 23 | 杜静.冠突散囊菌对中药材三七成分的转化及机理研究[D].贵阳:贵州师范大学,2020. |
| DU J. Biotransformation of bioactive compounds in Pannax notoginseng fermented by Eurotium cristatum and its mechanism [D]. Guiyang: Guizhou Normal University, 2020. | |
| 24 | 王琪琪.黑茶中散囊属真菌及其对茶叶品质提升研究[D].贵阳:贵州师范大学,2021. |
| WANG Q Q. The diversity of Eurotium sp . in dark tea and its effect on the quality improvement [D]. Guiyang: Guizhou Normal University, 2021. | |
| 25 | REN X X, WANG Y C, LIU Y X, et al.. Comparative transcriptome analysis of the calcium signaling and expression analysis of sodium/calcium exchanger in Aspergillus cristatus [J]. J. Basic. Microbiol., 2018, 58(1):76-87. |
| 26 | WANG D Y, TONG S M, GUAN Y Y, et al.. The velvet protein vea functions in asexual cycle, stress tolerance and transcriptional regulation of Beauveria bassiana [J]. Fungal. Genet. Biol., 2019, 127:1-11. |
| 27 | LIU H, SANG S L, WANG H, et al.. Vea comparative proteomic analysis reveals the regulatory network of the gene during asexual and sexual spore development of Aspergillus cristatus [J/OL]. Biosci. Rep., 2018, 38(4):67 [2022-02-10]. . |
| 28 | 张玉婷,李伟国,梁冬梅,等. P450s在萜类合成方面的合成生物学研究进展[J].中国生物工程杂志,2020,40(8):84-96. |
| ZHANG Y T, LI W G, LIANG D M, et al.. Research progress in synthetic biology of P450s in terpenoid synthesis [J]. Chin. J. Biotechnol., 2020, 40(8):84-96. | |
| 29 | CHANG M, KEASLING J. Production of isoprenoid pharmaceuticals by engineered microbes [J]. Nat. Chem. Biol., 2006, 2(12):674-681. |
| 30 | CARY J W, HAN Z, YIN Y, et al.. Transcriptome analysis of Aspergillus flavus reveals vea-dependent regulation of secondary metabolite gene clusters, including the novel aflavarin cluster [J]. Eukaryot. Cell., 2015, 14(10):983-997. |
| 31 | 张梦薇.过表达laeA和veA基因对黑曲霉生长和代谢的影响[D].天津:天津科技大学,2020. |
| ZHANG M W. Effects of overexpression of laeA and veA genes on growth and metabolism of Aspergillus niger [D]. Tianjin: Tianjin University of Science and Technology, 2020. | |
| 32 | KONG W P, HUANG C S, SHI J, et al.. Recycling of Chinese herb residues by endophytic and probiotic fungus Aspergillus cristatus Cb10002 for the production of medicinal valuable anthraquinones [J/OL]. Microb. Cell. Fact., 2019, 18(1):9 [2022-02-10]. . |
| [1] | Yuqing ZHOU, Yongfei YANG, Changwei GE, Qian SHEN, Siping ZHANG, Shaodong LIU, Huijuan MA, Jing CHEN, Ruihua LIU, Shicong LI, Xinhua ZHAO, Cundong LI, Chaoyou PANG. Identification of Cold-related Co-expression Modules in Cotton Cotyledon by WGCNA [J]. Journal of Agricultural Science and Technology, 2022, 24(4): 52-62. |
| [2] | LI Shuxin, ZHANG Hao, ZHENG Housheng, ZHENG Peihe, PANG Shifeng, XU Shiquan. Transcriptome Analysis of Phenotypic Differences Between Ermaya and Changbo of Forest Cultivated Panax ginseng [J]. Journal of Agricultural Science and Technology, 2021, 23(9): 56-68. |
| [3] | LIU Yuan, ZHANG Xiuyan, XU Miaoyun, ZHENG Hongyan, ZOU Junjie, ZHANG Lan, WANG Lei. Global Small RNA Transcriptome Profiling of Rice Under Drought Stress [J]. Journal of Agricultural Science and Technology, 2021, 23(6): 23-32. |
| [4] | ZHANG Wenyun1, ZHANG Jiancheng2, YAO Jingzhen2*. Comparative Transcriptome Analysis of Wheat Leaf in Response to Low Nitrogen Stress#br# [J]. Journal of Agricultural Science and Technology, 2020, 22(11): 26-34. |
| [5] | CHEN Hailong1, JIANG Lihua1, CHEN Shuai1, ZHANG Xiaoge1, ZHU Nianqing1, WEI Pinghe1*, ZHOU Changlin2. Adk1 Overexpression and Sodium Citrate Feeding Enhanced S-adenosylmethionine Synthesis in Yeast [J]. Journal of Agricultural Science and Technology, 2020, 22(10): 69-76. |
| [6] | CHEN Cheng1, LIU Xiaofei2, LI Qiang3, WANG Jian1, FU Rongtao1, ZHANG Hong1, LU Daihua1*. Bioinformatic Analysis of Simple Sequence Repeat (SSR) Loci in Ganoderma oregonense Transcriptome [J]. Journal of Agricultural Science and Technology, 2018, 20(7): 48-55. |
| [7] | WANG Lin1, YANG Lijun2, ZHAO Jiajia2, HUANG Haitang2, XU Zicheng1*. Application of Omics Analysis Technology in Tobacco Research [J]. Journal of Agricultural Science and Technology, 2018, 20(7): 56-62. |
| [8] | HUANG Juan, DENG Jiao, CHEN Qingfu*. Transcriptome Analysis of Fagopyrum Root and Identification of Genes Involved in Flavonoid Biosynthesis [J]. Journal of Agricultural Science and Technology, 2017, 19(2): 9-19. |
| [9] | YUE Chun-jiang1§, CHEN Chuan-chuan1§, GUO Feng-xian1, LI Hua1, SUN Hong-bo1, PEI Dan-ning1, MA Xiao-qing1, CHEN Fu-xin1, YANG Huo-li1, LI Qin1, LIU Yue1,2*. Data Mining of Simple Sequence Repeats in Transcriptome Sequences of Mongolia Medicinal Plant Artemisia frigida Willd [J]. Journal of Agricultural Science and Technology, 2016, 18(6): 31-43. |
| [10] | LI Rui-yuan1,PAN Fan2, CHEN Qing-fu2, SHI Tao-xiong2*. Excavation and Polymorphism Analysis of EST-SSR from Transcriptome of Tartary Buckwheat [J]. , 2015, 17(4): 42-52. |
| [11] | SU Shiyou, TENG Chao, ZHANG Wei, CHEN Ming*. Progress on Type Ⅲ Polyketide Synthase from Bacteria [J]. , 2013, 15(6): 119-129. |
| [12] | WU Mao-sen, LI Lei, WU Jing, CHEN Hua-min, HE Chen-yang. Effects of Overexpression and RNAi-mediated Silence of OsCtBP-A Gene on Root Growth and Stress Resistance in Rice [J]. , 2010, 12(1): 106-110. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
 京公网安备11010802021197号
京公网安备11010802021197号