




















中国农业科技导报 ›› 2024, Vol. 26 ›› Issue (1): 78-88.DOI: 10.13304/j.nykjdb.2022.0622
李相吴1,2( ), 刘自扬2,3, 徐玉俊2, 祝建波1(
), 刘自扬2,3, 徐玉俊2, 祝建波1( ), 吴燕民2,3(
), 吴燕民2,3( )
)
收稿日期:2022-07-23
接受日期:2022-09-28
出版日期:2024-01-15
发布日期:2024-01-08
通讯作者:
祝建波,吴燕民
作者简介:李相吴 E-mail:1003786451@qq.com;
基金资助:
Xiangwu LI1,2( ), Ziyang LIU2,3, Yujun XU2, Jianbo ZHU1(
), Ziyang LIU2,3, Yujun XU2, Jianbo ZHU1( ), Yanmin WU2,3(
), Yanmin WU2,3( )
)
Received:2022-07-23
Accepted:2022-09-28
Online:2024-01-15
Published:2024-01-08
Contact:
Jianbo ZHU,Yanmin WU
摘要:
为探讨真菌诱导子调控紫草素合成的分子机制,以新疆紫草无菌苗为试材,经尖孢镰刀菌、立枯丝核菌制备成的诱导子生物诱导后,对其根部进行RNA测序和分析。结果表明,与对照组比较,尖孢镰刀菌试验组差异表达基因1 735个;立枯丝核菌试验组差异表达基因1 043个。GO(gene ontology)分析发现,2个试验组差异表达基因主要富集在细胞过程、细胞组分中膜和分子功能中催化活性等生物学过程。KEGG(kyoto encyclopedia of genes and genomes)分析发现,2个试验组在植物与病原菌互作、植物激素信号转导途径和苯丙素合成等通路均有大量差异表达基因富集。2个试验组差异表达情况相同的转录因子主要包括bHLH、AP2/ERF-ERF和LOB等,并发现参与紫草素合成及其正向调控的AeGHQH、AeDSH1、AeAP、AePAL、AeDI2、AePGT、AeHMGR、AeG10H基因在2个试验组均上调表达,其中尖孢镰刀菌试验组上调表达较显著。以上结果从分子水平探究了新疆紫草对真菌诱导子生物诱导的响应机制,为未来真菌诱导子应用于新疆紫草种植生产奠定了理论基础。
中图分类号:
李相吴, 刘自扬, 徐玉俊, 祝建波, 吴燕民. 真菌诱导子调控紫草素合成的分子机制探究[J]. 中国农业科技导报, 2024, 26(1): 78-88.
Xiangwu LI, Ziyang LIU, Yujun XU, Jianbo ZHU, Yanmin WU. Explore of Molecular Mechanism on Fungal Elicitors Regulating Shikonin Synthesis[J]. Journal of Agricultural Science and Technology, 2024, 26(1): 78-88.
处理 Treatment | 样品编号 Sample ID | 总碱基数 Base number | GC含量 GC content/% | Q30/% |
|---|---|---|---|---|
| F20 | F20-1 | 6 293 089 734 | 43.61 | 92.39 |
| F20-2 | 6 422 595 670 | 43.56 | 93.06 | |
| F20-3 | 7 834 038 640 | 43.62 | 92.46 | |
| R20 | R20-1 | 6 872 640 242 | 43.61 | 93.02 |
| R20-2 | 7 140 087 048 | 43.71 | 93.05 | |
| R20-3 | 6 946 752 472 | 43.66 | 92.70 | |
| CK | CK-1 | 7 792 349 390 | 43.66 | 92.82 |
| CK-2 | 6 870 409 240 | 43.68 | 92.81 | |
| CK-3 | 5 930 234 288 | 43.68 | 92.44 |
表1 样品测序数据评估
Table 1 sample sequencing data evaluation
处理 Treatment | 样品编号 Sample ID | 总碱基数 Base number | GC含量 GC content/% | Q30/% |
|---|---|---|---|---|
| F20 | F20-1 | 6 293 089 734 | 43.61 | 92.39 |
| F20-2 | 6 422 595 670 | 43.56 | 93.06 | |
| F20-3 | 7 834 038 640 | 43.62 | 92.46 | |
| R20 | R20-1 | 6 872 640 242 | 43.61 | 93.02 |
| R20-2 | 7 140 087 048 | 43.71 | 93.05 | |
| R20-3 | 6 946 752 472 | 43.66 | 92.70 | |
| CK | CK-1 | 7 792 349 390 | 43.66 | 92.82 |
| CK-2 | 6 870 409 240 | 43.68 | 92.81 | |
| CK-3 | 5 930 234 288 | 43.68 | 92.44 |
处理 Treatment | 样品编号 Sample ID | Read数 Reads number | 匹配Read数 Mapped reads number | 匹配率 Mapped ratio/% |
|---|---|---|---|---|
| F20 | F20-1 | 21 096 427 | 15 558 872 | 73.75 |
| F20-2 | 21 504 765 | 16 195 509 | 75.31 | |
| F20-3 | 26 236 242 | 19 430 135 | 74.06 | |
| R20 | R20-1 | 23 005 518 | 17 277 331 | 75.10 |
| R20-2 | 23 913 025 | 17 644 691 | 73.79 | |
| R20-3 | 23 283 171 | 17 129 464 | 73.57 | |
| CK | CK-1 | 26 093 101 | 19 723 787 | 75.59 |
| CK-2 | 23 036 789 | 17 376 776 | 75.43 | |
| CK-3 | 19 879 659 | 14 903 996 | 74.97 |
表2 测序数据与组装结果的比对
Table 2 Comparison of sequencing data with assembly results
处理 Treatment | 样品编号 Sample ID | Read数 Reads number | 匹配Read数 Mapped reads number | 匹配率 Mapped ratio/% |
|---|---|---|---|---|
| F20 | F20-1 | 21 096 427 | 15 558 872 | 73.75 |
| F20-2 | 21 504 765 | 16 195 509 | 75.31 | |
| F20-3 | 26 236 242 | 19 430 135 | 74.06 | |
| R20 | R20-1 | 23 005 518 | 17 277 331 | 75.10 |
| R20-2 | 23 913 025 | 17 644 691 | 73.79 | |
| R20-3 | 23 283 171 | 17 129 464 | 73.57 | |
| CK | CK-1 | 26 093 101 | 19 723 787 | 75.59 |
| CK-2 | 23 036 789 | 17 376 776 | 75.43 | |
| CK-3 | 19 879 659 | 14 903 996 | 74.97 |
| 数据库名称Database name | 被注释到的差异基因数量 Annotated number of DGEs | |
|---|---|---|
| F20/CK | R20/CK | |
| COG | 517 | 317 |
| GO | 1 276 | 785 |
| KEGG | 1 063 | 651 |
| KOG | 758 | 439 |
| Pfam | 1 294 | 775 |
| Swiss-Prot | 1 188 | 731 |
| NR | 1 555 | 943 |
表3 差异基因功能注释统计
Table 3 Numbers of functional annotated DEGs
| 数据库名称Database name | 被注释到的差异基因数量 Annotated number of DGEs | |
|---|---|---|
| F20/CK | R20/CK | |
| COG | 517 | 317 |
| GO | 1 276 | 785 |
| KEGG | 1 063 | 651 |
| KOG | 758 | 439 |
| Pfam | 1 294 | 775 |
| Swiss-Prot | 1 188 | 731 |
| NR | 1 555 | 943 |
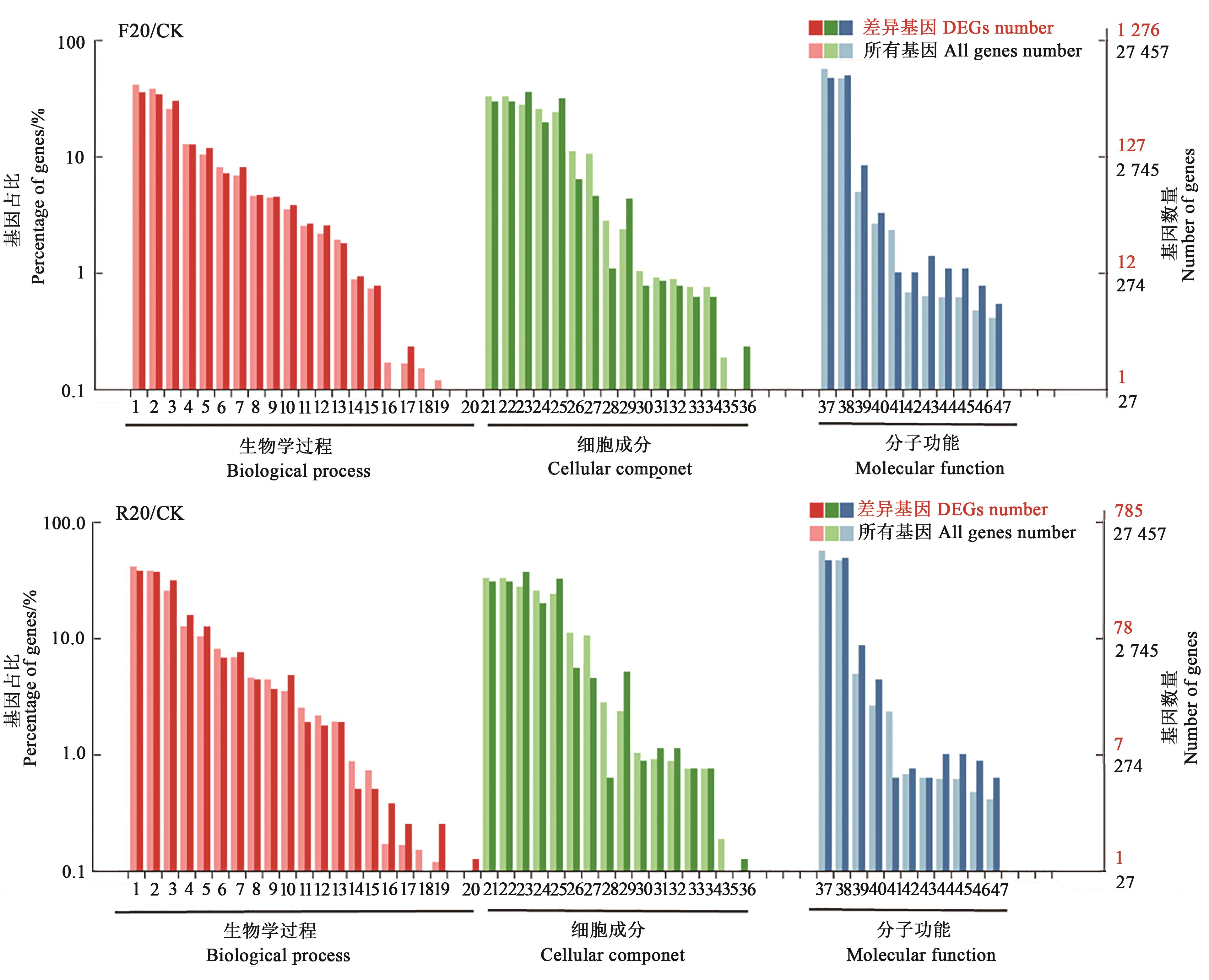
图2 差异基因GO功能注释注释:1—细胞过程;2—代谢过程;3—单一有机体过程;4—生物调节;5—刺激反应;6—细胞成分组织或生物合成;7—定位;8—发育过程;9—多细胞生物过程;10—信号;11—再生过程;12—多生物过程;13—再生;14—生长;15—免疫系统过程;16—节律过程;17—细胞运动;18—生物附着;19:解毒作用;20细胞聚合;21—细胞;22—细胞组分;23—细胞膜;24—细胞器;25—膜成分;26—细胞器组分;28—膜封闭腔;29—胞外区;30—胞外区组分;31—细胞连接;32—共生体;33—其他有机体;34—其他有机体成分;35—超分子复合物;36—内核;37—结合;38—催化活动;39—转运活动;40—核酸结合转录因子活性;41—结构分子活性;42—电子载体活性;43—抗氧化活动;44—分子转导活性;45—信号转导活性;46—转录因子活性,蛋白结合;47—分子功能调控。
Fig. 2 GO functional annotation of differential expressed genesNote:1—Cellular process; 2—Metabolic process; 3—Single-organism process; 4—Biological regulation; 5—Response to stimulus; 6—Cellular component organization or biogenesis; 7—Localization; 8—Developmental process; 9—Multicellular organismal process; 10—Signaling; 11—Reproductive process; 12—Multi-organism process; 13—Reproduction; 14—Growth; 15—Immune system process; 16—Rhythmic process; 17—Locomotion; 18—Biological adhesion; 19—Detoxification; 20—Cell aggregation; 21—Cell; 22—Cell part; 23—membrane; 24—Organelle; 25—Membrane part; 26—Organelle part; 27—Macromolecular complex; 28—Membrane-enclosed lumen; 29—Extracellular region; 30—Extracellular region part; 31—Cell junction; 32—Symplast; 33—Other organism; 34—Other organism part; 35—Supramolecular complex; 36—Nucleoid; 37—Binding; 38—Catalytic activity; 39—Transporter activity; 40—Nucleic acid binding transcription factor activity; 41—Structural molecule activity; 42—Electron carrier activity; 43—Antioxidant activity; 44—Molecular transducer activity; 45—Signal transducer activity; 46—Transcription factor activity protein binding; 47—Molecular function regulator.
家族 Family | 基因ID Gene ID | log2FC | |
|---|---|---|---|
| F20/CK | R20/CK | ||
| TRAF | c65742.graph_c0 | 2.51 | 2.01 |
| Tify | c54302.graph_c0 | 1.59 | 0.96 |
| SRS | c54737.graph_c1 | -4.53 | -2.95 |
| MYB-related | c58018.graph_c0 | 4.26 | 2.80 |
| MYB | c62808.graph_c0 | 1.59 | 1.15 |
| LOB | c48779.graph_c0 | 1.71 | 1.28 |
| HB-other | c47461.graph_c0 | -2.71 | -1.85 |
| HB-BELL | c58247.graph_c0 | 1.70 | 1.54 |
| GARP-G2-like | c65836.graph_c0 | -1.95 | -1.23 |
| GARP-G2-like | c53563.graph_c1 | -1.78 | -1.82 |
| C2H2 | c56397.graph_c0 | -2.06 | -1.52 |
| C2C2-YABBY | c28651.graph_c0 | -3.50 | -2.26 |
| C2C2-Dof | c53061.graph_c0 | 1.71 | 1.50 |
| bHLH | c59392.graph_c0 | 2.16 | 1.43 |
| bHLH | c54786.graph_c0 | 1.65 | 1.66 |
| bHLH | c53656.graph_c0 | -6.60 | -3.19 |
| bHLH | c47522.graph_c0 | -2.99 | -1.71 |
| AP2/ERF-ERF | c60825.graph_c0 | -1.78 | -1.17 |
| AP2/ERF-ERF | c57468.graph_c0 | 2.29 | 1.93 |
| AP2/ERF-ERF | c55718.graph_c0 | -1.68 | -1.33 |
表4 部分差异表达的转录因子(前20)
Table 4 Some differentially expressed transcription factor (top 20)
家族 Family | 基因ID Gene ID | log2FC | |
|---|---|---|---|
| F20/CK | R20/CK | ||
| TRAF | c65742.graph_c0 | 2.51 | 2.01 |
| Tify | c54302.graph_c0 | 1.59 | 0.96 |
| SRS | c54737.graph_c1 | -4.53 | -2.95 |
| MYB-related | c58018.graph_c0 | 4.26 | 2.80 |
| MYB | c62808.graph_c0 | 1.59 | 1.15 |
| LOB | c48779.graph_c0 | 1.71 | 1.28 |
| HB-other | c47461.graph_c0 | -2.71 | -1.85 |
| HB-BELL | c58247.graph_c0 | 1.70 | 1.54 |
| GARP-G2-like | c65836.graph_c0 | -1.95 | -1.23 |
| GARP-G2-like | c53563.graph_c1 | -1.78 | -1.82 |
| C2H2 | c56397.graph_c0 | -2.06 | -1.52 |
| C2C2-YABBY | c28651.graph_c0 | -3.50 | -2.26 |
| C2C2-Dof | c53061.graph_c0 | 1.71 | 1.50 |
| bHLH | c59392.graph_c0 | 2.16 | 1.43 |
| bHLH | c54786.graph_c0 | 1.65 | 1.66 |
| bHLH | c53656.graph_c0 | -6.60 | -3.19 |
| bHLH | c47522.graph_c0 | -2.99 | -1.71 |
| AP2/ERF-ERF | c60825.graph_c0 | -1.78 | -1.17 |
| AP2/ERF-ERF | c57468.graph_c0 | 2.29 | 1.93 |
| AP2/ERF-ERF | c55718.graph_c0 | -1.68 | -1.33 |
| 1 | 国家药典委员会.中华人民共和国药典-一部[M].北京:中国医药科技出版社,2020:355-356. |
| National Pharmacopoeia Committee. Pharmacopoeia of the People’s Republic of China-part1 [M]. Beijing: Chinese Medicine Science and Technology Press, 2020:355-356. | |
| 2 | WANG Y, ZHU Y, XIAO L, et al... Meroterpenoids isolated from Arnebia euchroma (Royle) Johnst. and their cytotoxic activity in human hepatocellular carcinoma cells [J]. Fitoterapia, 2018, 131(11):236-244. |
| 3 | CRUICKSHANK I, PERRIN D R. The isolation and partial characterization of monilicolin A, a polypeptide with phaseollin-inducing activity from Monilinia fructicola [J]. Life Sci., 1968, 7(10):449-458. |
| 4 | OKSMAN-CALDENTEY K, VERPOORTE R, VAN DER HEIJDEN R, et al... Engineering the plant cell factory for secondary metabolite production [J]. Transgenic Res., 2000, 9(4):323-343. |
| 5 | WANG Y, DAI C C, CAO J L, et al.. Comparison of the effects of fungal endophyte Gilmaniella sp. and its elicitor on atractylodes lancea plantlets [J]. World J. Microbiol. Biotechnol., 2011, 28(2):575-584. |
| 6 | ARGHAVANI P, HAGHBEEN K, MOUSAVI A. Enhancement of shikalkin production in Arnebia euchroma callus by a fungal elicitor, Rhizoctonia solani [J]. Iranian J. Biotechnol., 2015, 13(4):10-16. |
| 7 | 晏琼,胡宗定,吴建勇.生物与非生物诱导子协同作用对丹参毛状根培养生产丹参酮的影响[J].中国中药杂志,2006(3):188-191. |
| YAN Q, HU Z D, WU J Y. Synergistic effects of biotic and abiotic elicitors on the production of tanshinones in Salvia miltiorrhiza hairy root culture [J]. China J. Chin. Materia Medica, 2006(3):188-191. | |
| 8 | 田佩雯.白及内生真菌诱导子对宿主生长和主要活性物质的影响及调控[D].南宁:广西大学,2019. |
| TIAN P W. Effects and regulation of endophytic fungal elicitors from Bletilla striata on host growthand main substances [D]. Nanning: Guangxi University, 2019. | |
| 9 | 饶龙兵,杨汉波,郭洪英,等.不同倍性桤木属植物的转录组测序和分析[J].分子植物育种.2016,14(11):3047-3055. |
| RAO L B, YANG H B, GUO H Y, et al.. Analysis on transcriptome sequenced for alnus plants with different ploidy [J]. Mol. Plant Breed., 2016, 14(11):3047-3055. | |
| 10 | LOVE M I, HUBER W, ANDERS S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2 [J/OL]. Genome Biol., 2014, 15(12):550 [2022-06-02]. |
| 11 | GAISSER S, HEIDE L. Inhibition and regulation of shikonin biosynthesis in suspension cultures of Lithospermum [J]. Phytochemistry, 1996, 41(4):1065-1072. |
| 12 | SYKŁOWSKA-BARANEK K, PIETROSIUK A, NALIWAJSKI M R, et al.. Effect of L-phenylalanine on PAL activity and production of naphthoquinone pigments in suspension cultures of Arnebia euchroma (Royle) Johnst [J]. In Vitro Cellular Dev. Biol. Plant, 2012, 48(5):555-564. |
| 13 | YAZAKI K, KUNIHISA M, FUJISAKI T, et al.. Geranyl diphosphate:4-hydroxybenzoate geranyltransferase from Lithospermum erythrorhizon: cloning and characterization of a key enzyme in shikonin biosynthesis [J]. J. Biol. Chem., 2002, 277(8):6240-6246. |
| 14 | WANG S, WANG R S, LIU T G, et al.. CYP76B74 catalyzes the 3’’-hydroxylation of geranylhydroquinone in shikonin biosynthesis [J]. Plant Physiol. (Bethesda), 2019, 179(2):402-414. |
| 15 | 梁玖雯,李锬,王瑞杉,等.新疆紫草中2条CYP450基因的干扰毛状根体系的建立及其影响研究[J].中国中药杂志,2020,45(14):3422-3431. |
| LIANG J W, LI T, WANG R S, et al.. Establishment of RNA interfered hairy root system of two CYP450 genes in Arnebia euchroma and its influence [J]. China J. Chin. Materia Medica, 2020, 45(14):3422-3431. | |
| 16 | LIAO M, ZENG C, LIANG F. Two new dimeric naphthoquinones from Arnebia euchroma [J]. Phytochem. Letters, 2020, 37(1):106-109. |
| 17 | YAZAKI K, MATSUOKA H, SHIMOMURA K, et al.. A novel dark-inducible protein, LeDI-2, and its involvement in root-specific secondary metabolism in Lithospermum erythrorhizon [J]. Plant Physiol. (Bethesda), 2001, 125(3):1831-1841. |
| 18 | YAMAMURA Y N C U, SAHIN F P, NAGATSU A, et al.. Molecular cloning and characterization of a cDNA encoding a novel apoplastic protein preferentially expressed in a shikonin-producing callus strain of Lithospermum erythrorhizon [J]. Plant Cell Physiol., 2003, 44(4):437-446. |
| 19 | SAHA S, PAL D. Elicitor Signal Transduction Leading to the Production of Plant Secondary Metabolites [M]. Cham: Springer International Publishing, 2020:1-39. |
| 20 | 周雅涵.水杨酸、膜醭毕赤酵母、壳寡糖诱导柑橘果实抗病性及其生物学机制研究[D].重庆:西南大学,2017. |
| ZHOU Y H. Salicylic acid, Pichia membranaefaciens and oligochitosan induced disease resistance of citrus fruit and the possible biological mechanisms involved [D]. Chongqing: Southwest University, 2017. | |
| 21 | 瞿巾卓.酿酒葡萄细胞对内生真菌诱导子的代谢响应与机制[D].昆明:云南大学,2020. |
| JU J Z. Metabolic response and mechanism of Wine grape cells to elicitors from fungal endophytes [D]. Kunming: Yunnan University, 2020 | |
| 22 | 张明菊,朱莉,夏启中.植物激素对胁迫反应调控的研究进展[J].湖北大学学报(自然科学版).2021,43(3):242-253. |
| ZHANG M J, ZHU L, XIA Q ZAND. Research progress on the regulation of plant hormones to stress responses [J]. J. Hubei Univ., 2021, 43(3):242-253. | |
| 23 | SHAH L, ALI A, ZHU Y L, et al.. Wheat defense response to Fusarium head blight and possibilities of its improvement [J]. Physiol. Mol. Plant Pathol., 2017, 98(2):9-17. |
| 24 | NEMESIO-GORRIZ M, BLAIR P B, DALMAN K, et al.. Identification of Norway spruce MYB-bHLH-WDR transcription factor complex members linked to regulation of the flavonoid pathway [J/OL]. Front. Plant Sci., 2017, 8:305 [2022-06-02]. . |
| 25 | GRAEFF M, STRAUB D, EGUEN T, et al.. Microprotein-mediated recruitment of constans into a topless trimeric complex represses flowering in Arabidopsis [J/OL]. PLoS Genet., 2016, 12(3):e1005959 [2022-06-02]. . |
| 26 | DENG B, HUANG Z, GE F, et al.. An AP2/ERF family transcription factor PnERF1 raised the biosynthesis of saponins in panax notoginseng [J]. J. Plant Growth Regul., 2017, 36(3):691-701. |
| 27 | GAUTAM J K, GIRI M K, SINGH D, et al.. MYC2 influences salicylic acid biosynthesis and defense against bacterial pathogens in Arabidopsis thaliana [J]. Physiol. Plantarum., 2021, 173(4):2248-2261. |
| 28 | KAZAN K, MANNERS J M. MYC2: the master in action [J]. Mol. Plant, 2013, 6(3):686-703. |
| 29 | WANG F, ZHU H, CHEN D, et al.. A grape bHLH transcription factor gene, VvbHLH1, increases the accumulation of flavonoids and enhances salt and drought tolerance in transgenic Arabidopsis thaliana [J]. Plant Cell. Tissue Organ Cult., 2016, 125(2):387-398. |
| 30 | ZHANG M, LI S T, NIE L, et al.. Two jasmonate-responsive factors, TcERF12 and TcERF15, respectively act as repressor and activator of tasy gene of taxol biosynthesis in Taxus chinensis [J]. Plant Mol. Biol., 2015, 89(4-5):463-473. |
| 31 | EL-SAYED A S A, MOHAMED N Z, SAFAN S, et al.. Restoring the taxol biosynthetic machinery of aspergillus terreus by Podocarpus gracilior pilger microbiome, with retrieving the ribosome biogenesis proteins of WD40 superfamily [J]. Sci. Rep., 2019, 9(1):11512-11534. |
| 32 | HAN Z, YANG T, GUO Y, et al.. The transcription factor PagLBD3 contributes to the regulation of secondary growth in Populus [J]. J. Exp. Bot., 2021, 72(20):7092-7106. |
| 33 | TANG X M, WANG X, HUANG Y, et al... Natural variations of TFIIAγ gene and LOB1 promoter contribute to citrus canker disease resistance in Atalantia buxifolia [J]. PLoS Genetics. 2021, 17(1):e1009316 [2022-06-02]. . |
| 34 | HAO H, LEI C, DONG Q, et al... Effects of exogenous methyl jasmonate on the biosynthesis of shikonin derivatives in callus tissues of Arnebia euchroma [J]. Appl. Biochem. Biotechnol., 2014, 173(8):2198-2210. |
| 35 | WANG C G, WU J Y, MEI X G. Enhancement of taxol production and excretion in Taxus chinensis cell culture by fungal elicitation and medium renewal [J]. Appl. Microbiol. Biotechnol., 2001, 55(4):404-410. |
| [1] | 王潇然, 李笑语, 孙慧, 于海东, 石永春. 硼胁迫下烟草叶片转录组分析[J]. 中国农业科技导报, 2023, 25(8): 53-64. |
| [2] | 王云胜, 陈银翠, 程在, 张锦, 张传博. 过表达veA基因对冠突散囊菌次级代谢的影响[J]. 中国农业科技导报, 2023, 25(7): 77-86. |
| [3] | 马蓝, 彭晴, 徐小轻, 杨硕, 张宇微, 田丹丹, 施琳波, 石波, 乔宇. 大肠杆菌O157∶H7生物被膜状态下基因表达分析[J]. 中国农业科技导报, 2023, 25(6): 71-88. |
| [4] | 周雨青, 杨永飞, 葛常伟, 沈倩, 张思平, 刘绍东, 马慧娟, 陈静, 刘瑞华, 李士丛, 赵新华, 李存东, 庞朝友. 基于WGCNA的棉花子叶抗冷相关共表达模块鉴定[J]. 中国农业科技导报, 2022, 24(4): 52-62. |
| [5] | 郭瑞锋, 任月梅, 杨忠, 刘贵山, 任广兵, 张绶, 朱文娟. 草甘膦铵盐诱导谷子雄性不育的转录组分析[J]. 中国农业科技导报, 2022, 24(10): 35-43. |
| [6] | 李舒欣, 张浩, 郑厚胜, 郑培和, 逄世峰, 许世泉. 转录组分析二马牙和长脖类型林下参表型差异[J]. 中国农业科技导报, 2021, 23(9): 56-68. |
| [7] | 刘源, 张秀妍, 徐妙云, 郑红艳, 邹俊杰, 张兰, 王磊. 水稻干旱胁迫的small RNA转录组分析[J]. 中国农业科技导报, 2021, 23(6): 23-32. |
| [8] | 张文云1,张建诚2,姚景珍2*. 氮胁迫下小麦叶片转录组分析[J]. 中国农业科技导报, 2020, 22(11): 26-34. |
| [9] | 陈诚1,刘晓飞2,李强3,王剑1,伏荣桃1,张鸿1,卢代华1*. 美国大灵芝(Ganoderma oregonense)转录组SSR位点的生物信息学分析[J]. 中国农业科技导报, 2018, 20(7): 48-55. |
| [10] | 王林1,杨立均2,赵佳佳2,黄海棠2,许自成1*. 组学分析技术在烟草研究中的应用进展[J]. 中国农业科技导报, 2018, 20(7): 56-62. |
| [11] | 黄娟,邓娇,陈庆富*. 荞麦根的转录组学分析及黄酮合成基因的鉴定[J]. 中国农业科技导报, 2017, 19(2): 9-19. |
| [12] | 岳春江1§,陈川川1§,郭凤仙1,李华1,孙洪波1,裴丹宁1,马晓清1,陈富欣1,杨获莉1,李琴1,刘越1,2*. 蒙药冷蒿转录组SSR信息分析[J]. 中国农业科技导报, 2016, 18(6): 31-43. |
| [13] | 黎瑞源1,潘凡2,陈庆富2,石桃雄2*. 苦荞转录组EST-SSR发掘及多态性分析[J]. , 2015, 17(4): 42-52. |
| [14] | 王辰,赵为民,牟玉莲*. 调控动物表型microRNA在分子育种中新的重要标记[J]. , 2013, 15(3): 108-112. |
| [15] | 李晓晖,李鑫鑫,张维,燕永亮,陈明,陆伟. 宏转录组学在微生物生态学研究中的应用[J]. , 2011, 13(4): 58-65. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
 京公网安备11010802021197号
京公网安备11010802021197号